|
|
Corces-Zimmerman, M. R., & Majeti, R. (2014). Pre-leukemic evolution of hematopoietic stem cells: the importance of early mutations in leukemogenesis. Leukemia, 28(12), 2276–2282.

Abstract: Cancer has been shown to result from the sequential acquisition of genetic alterations in a single lineage of cells. In leukemia, increasing evidence has supported the idea that this accumulation of mutations occurs in self-renewing hematopoietic stem cells (HSCs). These HSCs containing some, but not all, leukemia-specific mutations have been termed as pre-leukemic. Multiple recent studies have sought to understand these pre-leukemic HSCs and determine to what extent they contribute to leukemogenesis. These studies have elucidated patterns in mutation acquisition in leukemia, demonstrated resistance of pre-leukemic cells to standard induction chemotherapy and identified these pre-leukemic cells as a putative reservoir for the generation of relapsed disease. When combined with decades of research on clonal evolution in leukemia, mouse models of leukemogenesis, and recent massively parallel sequencing-based studies of primary patient leukemia, studies of pre-leukemic HSCs begin to piece together the evolutionary puzzle of leukemogenesis. These results have broad implications for leukemia treatment, targeted therapies, minimal residual disease monitoring and early detection screening.
|
|
|
|
Hauer, J., Borkhardt, A., Sanchez-Garcia, I., & Cobaleda, C. (2014). Genetically engineered mouse models of human B-cell precursor leukemias. Cell Cycle, 13(18), 2836–2846.

Abstract: B-cell precursor acute lymphoblastic leukemias (pB-ALLs) are the most frequent type of malignancies of the childhood, and also affect an important proportion of adult patients. In spite of their apparent homogeneity, pB-ALL comprises a group of diseases very different both clinically and pathologically, and with very diverse outcomes as a consequence of their biology, and underlying molecular alterations. Their understanding (as a prerequisite for their cure) will require a sustained multidisciplinary effort from professionals coming from many different fields. Among all the available tools for pB-ALL research, the use of animal models stands, as of today, as the most powerful approach, not only for the understanding of the origin and evolution of the disease, but also for the development of new therapies. In this review we go over the most relevant (historically, technically or biologically) genetically engineered mouse models (GEMMs) of human pB-ALLs that have been generated over the last 20 years. Our final aim is to outline the most relevant guidelines that should be followed to generate an “ideal” animal model that could become a standard for the study of human pB-ALL leukemia, and which could be shared among research groups and drug development companies in order to unify criteria for studies like drug testing, analysis of the influence of environmental risk factors, or studying the role of both low-penetrance mutations and cancer susceptibility alterations.
Keywords: B-precursor leukemia; Bcr-Abl; CLP, common lymphoid progenitor; GEMM, genetically engineered mouse model; LIC, leukemia-initiating cell; Mll; ROS, reactive oxygen species.; Tel-Aml1; mouse models; pB-ALL; pB-ALL, preB-Acute lymphoblastic leukemia
|
|
|
|
Gawad, C., Koh, W., & Quake, S. R. (2014). Dissecting the clonal origins of childhood acute lymphoblastic leukemia by single-cell genomics. Proc Natl Acad Sci U S A, 111(50), 17947–17952.

Abstract: Many cancers have substantial genomic heterogeneity within a given tumor, and to fully understand that diversity requires the ability to perform single cell analysis. We performed targeted sequencing of a panel of single nucleotide variants (SNVs), deletions, and IgH sequences in 1,479 single tumor cells from six acute lymphoblastic leukemia (ALL) patients. By accurately segregating groups of cooccurring mutations into distinct clonal populations, we identified codominant clones in the majority of patients. Evaluation of intraclonal mutation patterns identified clone-specific punctuated cytosine mutagenesis events, showed that most structural variants are acquired before SNVs, determined that KRAS mutations occur late in disease development but are not sufficient for clonal dominance, and identified clones within the same patient that are arrested at varied stages in B-cell development. Taken together, these data order the sequence of genetic events that underlie childhood ALL and provide a framework for understanding the development of the disease at single-cell resolution.
Keywords: acute lymphoblastic leukemia; clonal evolution; cytosine mutagenesis; intratumor heterogeneity; single-cell genomics
|
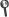
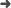
|
|
|
Alpar, D., Wren, D., Ermini, L., Mansur, M. B., van Delft, F. W., Bateman, C. M., et al. (2014). Clonal origins of ETV6-RUNX1 acute lymphoblastic leukemia: studies in monozygotic twins. Leukemia, .

Abstract: Studies on twins with concordant acute lymphoblastic leukemia (ALL) have revealed that ETV6-RUNX1 gene fusion is a common, pre-natal genetic event with other driver aberrations occurring subclonally and probably post-natally. The fetal cell type that is transformed by ETV6-RUNX1 is not identified by such studies or by the analysis of early B-cell lineage phenotype of derived progeny. Ongoing, clonal immunoglobulin (IG) and cross-lineage T-cell receptor (TCR) gene rearrangements are features of B-cell precursor leukemia and commence at the pro-B-cell stage of normal B-cell lineage development. We reasoned that shared clonal rearrangements of IG or TCR genes by concordant ALL in twins would be informative about the fetal cell type in which clonal advantage is elicited by ETV6-RUNX1. Five pairs of twins were analyzed for all varieties of IG and TCR gene rearrangements. All pairs showed identical incomplete or complete V(D)J junctions coupled with substantial, subclonal and divergent rearrangements. This pattern was endorsed by single cell genetic scrutiny in one twin pair. Our data suggest that the pre-leukemic initiating function of ETV6-RUNX1 fusion is associated with clonal expansion early in the fetal B-cell lineage.Leukemia accepted article preview online, 12 November 2014. doi:10.1038/leu.2014.322.
|
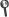
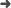
|
|
|
Yoon, H. E., Lee, J. S., Myung, S. H., & Lee, Y. - S. (2014). Increased γ-H2AX by exposure to a 60-Hz magnetic fields combined with ionizing radiation, but not hydrogen peroxide, in non-tumorigenic human cell lines. International journal of radiation biology, 90(4), 291–298.

Abstract: PURPOSE: Genotoxic effects have been considered the gold standard to determine if an environmental factor is a carcinogen, but the currently available data for extremely low frequency time-varying magnetic fields (ELF-MF) remain controversial. As an environmental stimulus, the effect of ELF-MF on cellular DNA may be subtle. Therefore, a more sensitive method and systematic research strategy are warranted to evaluate genotoxicity. MATERIALS AND METHODS: We investigated the effect of ELF-MF in combination with ionizing radiation (IR) or H(2)O(2) on the DNA damage response of expression of phosphorylated H2AX (γ-H2AX) and production of γ-H2AX foci in non-tumorigenic human cell systems consisting of human lung fibroblast WI-38 cells and human lung epithelial L132 cells. RESULTS: Exposure to a 60-Hz, 2 mT ELF-MF for 6 h produced increased γ-H2AX expression, as well as γ-H2AX foci production, a common DNA double-strand break (DSB) marker. However, exposure to a 1 mT ELF-MF did not have the same effect. Moreover, 2 mT ELF-MF exposure potentiated the expression of γ-H2AX and γ-H2AX foci production when combined with IR, but not when combined with H(2)O(2). CONCLUSIONS: ELF-MF could affect the DNA damage response and, in combination with different stimuli, provide different effects on γ-H2AX.
Keywords: Cells; Cultured; DNA Damage; Histones; Histones: metabolism; Humans; Hydrogen Peroxide; Hydrogen Peroxide: pharmacology; Ionizing; Magnetic Fields; Radiation; Sister Chromatid Exchange; Sister Chromatid Exchange: radiation effects
|
|

